|

|
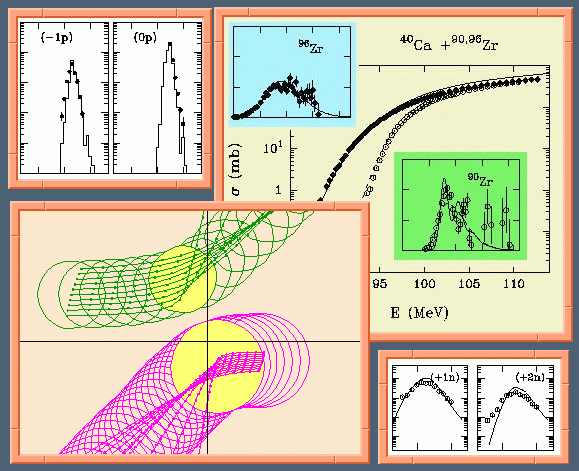
GRAZING_9 is an interactive Fortran program that evaluates the reactions to be expected in a collision between two heavy nuclei at moderate bombarding energies. It calculates the cross section for the distribution in mass and charge, kinetic energy and excitation energy of the two scattered nuclei. It also calculates the cross section leading to capture including coupled channel effect in the tunneling of the fusion barrier. It uses the NAG library Mark 18.
Here you find the zipped tar file that contains the executable of the
GRAZING program that should run on any
LINUX (there is a version at 32bits and one at 64bits).
To install unzip and untar the file in any location of your choice, there
you find the grazing_9 (the exe files at 32bits and the one at 64bits)
and a directory data containing all the file needed by GRAZING.
To run the program you MUST first define the enviromental variable
GRAZING_DIR that refer to the data directory. In the bash shell:
(Place these lines in .bash_profile) GRAZING_DIR='my_grazing_dir/data' export GRAZING_DIRLet me know if it does not wotk
Here you find the zipped tar file that contains the program: grazing_9 (393 Kb - June 2002). Notice this is the last version with quite few upgrades done by myself so the mistakes rest on me and not on the author. If you have problems please do not esitate to contact me (see at the bottom).
This distribution of GRAZING_9 has been tested to run both on
LINUX (redhat 6.2) and on ALPHA (Compaq TRU64-UNIX). The program uses the
NAG library (Mark 18).
Copy the .gz file in the directory you want to place grazing_9. Then
issue the following commands to unzip and untar the file:
>gunzip grazing.tar.gz
>tar xvf grazing.tar
After these operations you have a new directory grazing that contains all
the relevant files for grazing_9 and three (3) subdirectories:
grazing/doc contains some documentation about the program
grazing/pg contains the plotting program gzp2. You may find it
usefull, otherwise just discart it.
grazing/examples contains few scripts for running grazing_9. There is
also e file .inp for plotting with gzp2.
Now follows these steps to satisfy the local setting and the architecture
you are interested in:
LINUX - ALPHA
1 - grazing_file.icl
modify the file according to you local setting (use the full path name).
The files referred contain informations about exces masses of the nuclei
and the energy and strenght of the low lying states
2 - makefile
modify the file to satisfy your local setting for the NAG library
and other global definitions like compiler and the use of other
routines. Notise that these corrections appear at the beginning of
the file and at the end.
3 - issue the command
>make
If everithing is correct you should have the file grazing_9 that
contains the executable.
4 - Make sure you have installed on your machine the graphical PGPLOT
library. Go to the subdirectory pg. Modify the script flpg to satisfy
yout local setting and then issue the command:
>flpg gzp2
You should have the plotting program gzp2
5 - Go to the subdirectory examples, modify the file bd_com
and fus_com to satisfy your local setting
6 - Execute both scripts. You should have two new files .dat containing the
fusion cross sections and the barrier distribution
7 - issue the following command
>../pg/gzp2
pg_fus.inp
/XWINDOW
a new window should appear on the screen with the
excitation function and the barrier distribution you just calculated.
To close the window pres a C_RET from the command screen.
If you feel satisfied with gzp2 you may learn more about it consulting
the page: http:/www.to.infn.it/~nanni/gzp2.html.